As promising cancer treatments emerge the need for improved detection and characterization methods are still evident. Identification of novel biomarkers is a promising area of cancer research and development but because of the high complexity and heterogeneity of tumors much remains to be learned.
What is a cancer biomarker? A biomarker is a biological molecule that can be found in the blood, bodily fluids or tissue of interest (i.e. tumor) that can give information about the molecular characteristics of a tumor.
Specimens for biomarker discovery
- Tumor biopsy
- Blood
Potential biomarker biological molecules
- DNA (copy number, methylation states, mutations)
- RNA (mRNA, microRNA)
- Protein (phosphorylation, post-translational modifications)
- Metabolic products
Tools for cancer biomarker identification
- Genome mutation screening
- Gene expression profiling
- Protein and metabolomic expression profiling
Biomarkers can be used as tools for diagnosis (detect the presence of cancer), prognosis, tracking cancer progression, and assessing treatment efficacy.
In cancer, a biomarker is often a protein that is mutated or is expressed at higher levels in the cancerous cells compared to the normal tissue. There are various proteins whose mutated status is shared by multiple types of cancers these include inactivating mutations of tumor suppressor proteins such as the cell cycle regulators p53, PTEN, and retinoblastoma protein (RB) and activating mutations of proto-oncogenes such as Ras and Myc. Another prominent cancer biomarker is the cell proliferation marker, Ki67 that can be used not only as a prognostic indicator but also to assess the efficacy of a treatment where reduction in Ki67 expression indicates reduced cellular proliferation.
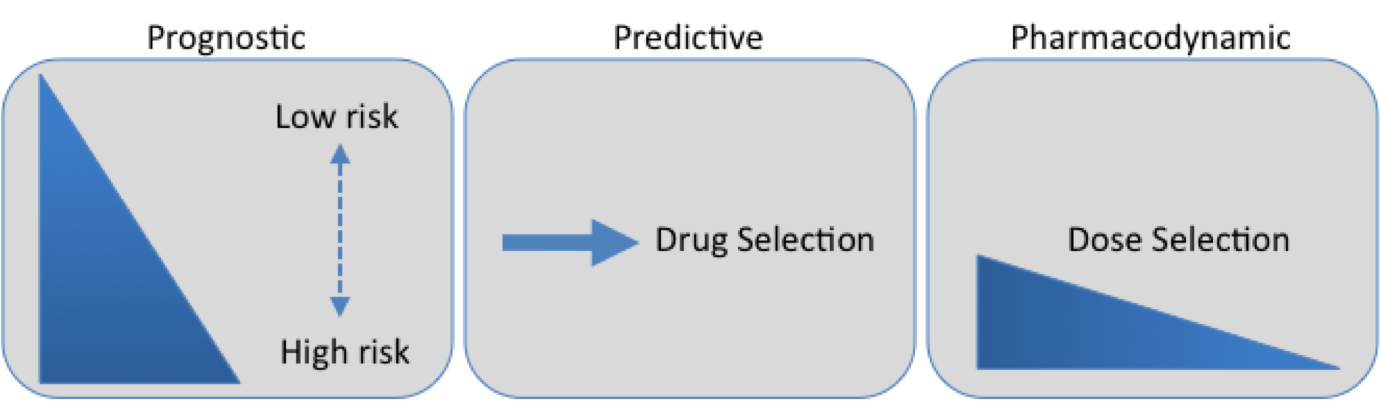
Types of biomarkers and their uses
- Prognostic biomarker: Knowing the key molecular changes in a patient’s cancer allows a doctor to determine whether the patient is likely to have a poor outcome and thus more aggressive treatment is necessary.
- Predictive biomarker: Understanding the molecular characteristics about a patients cancer can lead to tailoring of drug treatments with a higher likelihood of efficacy. For example, patients with certain kinase domain mutations on EGFR would possibly not respond to EGFR targeted treatments such as erlotinib. Additionally this gives the added benefit reducing a patients exposure to possible toxic side effects from a drug they may not have benefited from.
- Pharmacodynamic biomarker: Using biomarkers drug dosing could be tailored to each patient. Dosing a drug that has a specific molecular target can be decided based on its ability to decrease the activity of its biological target. For example, if a patient shows high activity for a particular kinase a targeted drug could be dosed up for that patients specific needs.
With the advent of genomic and proteomic technology and improved data mining algorithms, it is now easier and faster to identify biomarkers. Unfortunately, a major limitation in biomarker research and discovery is the need for biopsy samples and their limited availability for research. Another issue is that most biopsy samples available are taken from a patient during the initial diagnosis. Less available are samples from patients post treatment initiation or with advanced disease where the molecular characteristics of their cancer may have very likely changed.
Luckily, there is great interest in developing and improving current technologies to utilize blood specimens for protein, metabolic products, CTCs and circulating DNA as alternative non-invasive sources to identify and screen for cancer biomarkers.
Further reading:
Mining the plasma proteome for cancer biomarkers
Taming the dragon: genomic biomarkers to individualize the treatment of cancer
Proteomics for Cancer Biomarker Discovery