Forkhead box P3 (FoxP3)+ CD4+ T cells, known as regulatory T cells or TREGs, are a class of negative regulatory T cells that function to suppress immune responses, thereby establishing tolerance, preventing autoimmunity, and allowing tumor escapes from immune surveillance. TREGs are thought to be generated by two major mechanisms. Natural TREGs are generated through positive selection in the thymus via differential TCR signaling compared with conventional T cells. Adaptive or converted TREGs are thought be generated in the periphery by conversion of conventional CD4+ T cells via various mechanisms.
TREGs are a heterogeneous population of T cells that function via cell-contact dependent and independent mechanisms to suppress various immune cell types. Contact-dependent mechanisms of suppression include expression of negative regulatory receptors such as CTLA4, or killing of associated dendritic cells (DCs) through secretion of perforin and granzyme B. Contact-independant mechanisms of suppression include TREGs secretion of immune suppressive cytokines including IL-10 and TGFb. High expression of the IL-2 co-receptor CD25 allows TREGs to act as a sink for IL-2 thereby leading to IL-2 deprivation of conventional T cells and inhibition of proliferation.
TREGs are thus an important class of cells and study of these cell populations in human PBMC requires an understanding of the surface and intracellular markers that can be used for flow cytometry analysis and isolation by Fluorescence-activated cell sorting (FACS) or other methods.
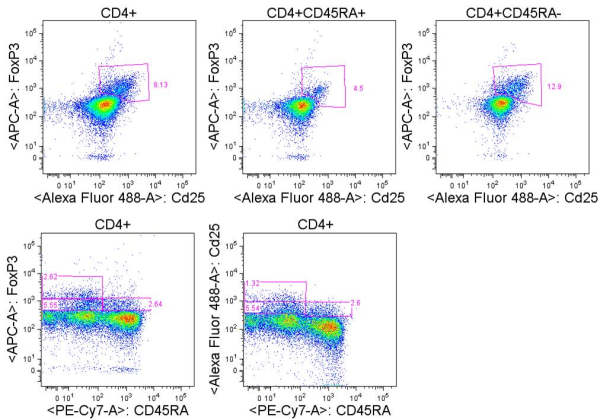
Miyara et. al. identified three functionally unique FoxP3+ populations in freshly isolated CD4+ T cells from human PBMC. These three populations could be identified by flow cytometry staining of CD45RA, FoxP3, and CD25. CD25 and FoxP3 expression were highly correlated in the CD4+ population, and I have consistently seen this in my own analyses of unstimulated human PBMC. The three populations included CD45RA+FoxP3low cells which were CD25++, CD45RA–FoxP3high cells which were CD25+++, and CD45RA–FoxP3low cells which were CD25 ++. When these populations were FACS sorted based on CD45RA and CD25 expression, only CD45RA+CD25++ and CD45RA–CD25+++ cells were functionally suppressive in co-culture experiments with TCR-activated CD25−CD45RA+CD4+ responder T cells. Thus CD45RA+FoxP3lowCD25++ cells and CD45RA–FoxP3highCD25+++ cells were denoted as naïve/resting and effector/activated TREGs, respectively. CD45RA–FoxP3low cells in contrast, are likely a heterogeneous mixture of cells and include some cells able to produce IFNg, IL-17, and IL-2 upon PMA+ ionomycin stimulation. Because dividing effector T cells are able to transiently express FoxP3 at low levels, these cells are likely to be contained in the CD45RA–FoxP3low population. Thus, when using CD25 or FoxP3 to identify TREGs by flow cytometry, CD45RA should be included, and care must be taken with the gating strategies.

CD25 in combination with TNFR2 and/or the lack of expression of CD127 have been shown to identify FoxP3+ TREGs that are highly suppressive even in CD25low populations and thus may be excellent markers in particular for FACS sorting of TREGs for functional analyses wherein FoxP3 cannot be utilized as a selection marker.
Several other markers have been used to delineate different populations of TREGs. The intracellular inhibitory receptor CTLA4, the co-stimulatory receptor ICOS, and the MHC class II cell surface receptor HLA-DR, are co-expressed with FoxP3 in the CD45RA–FoxP3high TREG population and may be utilized as specific markers of that population.
Depending on the assay conditions, additional markers may be used to identify TREGs. LAP, CD121a, and CD121b have been noted as highly specific markers of TREGs but are not expressed in the resting state, becoming transiently induced under assay conditions utilizing TCR stimulation.
This is by no means an exhaustive list of markers that have been used to identify human TREGs in their various functional subsets and states. The 2011 review in Int Immunopharmacol. by Chen et. al. discusses the usage of these and other markers including CCR6, LAG-3, GARP, CD103, CD39, and CD49d.
In summary, there are multiple combinations of markers that can be used to identify functionally different TREG populations within human PBMC. The selection of these markers should be considered in the context of the assay type being done and the questions being asked about these heterogeneous populations of cells.
Further Reading:
Regulatory T cells: mechanisms of differentiation and function. Josefowicz SZ, Lu LF, Rudensky AY. Annu Rev Immunol. 2012;30:531-64.
Foxp3+ regulatory T cells: differentiation, specification, subphenotypes. Feuerer M, Hill JA, Mathis D, Benoist C. Nat Immunol. 2009 Jul;10(7):689-95.
Functional delineation and differentiation dynamics of human CD4+ T cells expressing the FoxP3 transcription factor. Miyara M, Yoshioka Y, Kitoh A, Shima T, Wing K, Niwa A, Parizot C, Taflin C, Heike T, Valeyre D, Mathian A, Nakahata T, Yamaguchi T, Nomura T, Ono M, Amoura Z, Gorochov G, Sakaguchi S. Immunity. 2009 Jun 19;30(6):899-911.
Resolving the identity myth: key markers of functional CD4+FoxP3+ regulatory T cells. Chen X, Oppenheim JJ. Int Immunopharmacol. 2011 Oct;11(10):1489-96.
A peripheral circulating compartment of natural naive CD4 Tregs. D. Valmori, A. Merlo, N.E. Souleimanian, C.S. Hesdorffer, M. Ayyoub. J. Clin. Invest., 115 (2005), pp. 1953–1962.
Activation-induced FOXP3 in human T effector cells does not suppress proliferation or cytokine production. Allan SE, Crome SQ, Crellin NK, Passerini L, Steiner TS, Bacchetta R, Roncarolo MG, Levings MK. Int Immunol. 2007 Apr;19(4):345-54.
CD127 expression inversely correlates with FoxP3 and suppressive function of human CD4+ T reg cells. W. Liu, A.L. Putnam, Z. Xu-Yu, G.L. Szot, M.R. Lee, S. Zhu, P.A. Gottlieb, P. Kapranov, T.R. Gingeras, B. Fazekas de St Groth et al. J. Exp. Med., 203 (2006), pp. 1701–1711
Co-expression of TNFR2 and CD25 identifies more of the functional CD4+FOXP3+ regulatory T cells in human peripheral blood. Chen X, Subleski JJ, Hamano R, Howard OM, Wiltrout RH, Oppenheim JJ. Eur J Immunol. 2010 Apr;40(4):1099-106.