When cells undergo transformation and initiate the formation of a solid tumor mass, they cause profound changes on the phenotypes of the cells that surround them1. However, in addition to the changes in cellular phenotype, there is a change in the extracellular matrix that coincides with tumor formation1. It has been demonstrated that the majority of solid tumors have increased stiffness in their extracellular matrix (ECM), which may lead to increased activation of pro-tumor signaling pathways, such as Src, FAK, and RhoA2-4. Recently, it was discovered that increased matrix stiffness may also lead to increased activity of the oncogenic YAP/TAZ complex, which is connected to the Hippo signaling pathways, transcriptional regulators that increase cellular proliferation, decreased cellular contact inhibition, increased cancer stem cell phenotype, and increased metastasis5. However, in a  recent edition of Nature Cell Biology Calvo et al. demonstrated that 6. Not only do the authors demonstrate that YAP/TAZ is active in CAFs, but YAP/TAZ is necessary for CAF development6. They show that CAF activation leads to matrix remodeling towards increased stiffness, via myosin light chain 2 (MYL9/MLC) expression, establishing a feed-forward loop where the ECM plays a vital role6.
recent edition of Nature Cell Biology Calvo et al. demonstrated that 6. Not only do the authors demonstrate that YAP/TAZ is active in CAFs, but YAP/TAZ is necessary for CAF development6. They show that CAF activation leads to matrix remodeling towards increased stiffness, via myosin light chain 2 (MYL9/MLC) expression, establishing a feed-forward loop where the ECM plays a vital role6.
The authors first isolated fibroblasts in different stages towards becoming a CAF and saw that both mechanical-responsive signaling machinery (SMA, FN1, Paxillin, MYL9, MYH10, DIAH1 & F-actin) and mechanical tension were increased in populations containing CAFs. Moreover, tumor cell invasion, and angiogenesis of the tumor microenvironment (shown via endomucin and second-harmonic microscopy) were increased in samples that contained more tumor-associated-like fibroblasts (indicated by vimentin)6.
Because of the role of cell-cell and cell-ECM contact in the Hippo signaling pathway, the authors sought to understand whether this pathway is activated in CAFs. They found that YAP, and its co-factor, TAZ to be only upregulated and co-localized in the nucleus of transformed fibroblasts; the target of the activated YAP-TAZ complex6. Furthermore, upon depletion of YAP, the ability for CAFs to cause matrix stiffness by contraction lessened as well as CAFs ability to form collagen networks and facilitate angiogenesis. Interestingly, when TAZ was inhibited, there was no change in functionality, which may lead to a TAZ-independent function for YAP.
Upon microarray analysis of CAFs treated with siRNA that targets YAP, Calvo et al. found that the expression of many of the genes involved in mechanosensing and motility to be diminished6. Furthermore, when these individual genes were silenced, there was an overall decrease in the amount of cellular invasion of tumors. Many of the YAP-mediated genes, such as ANLN and DIAPH3, were involved in matrix remodeling and cellular invasion. Interestingly, modification of only one protein overexpression resulted in high amounts of matrix-remodeling and invasion: myosin regulatory light polypeptide 9 (MYL9). While not transcriptionally controlled by the YAP/TAZ complex, the authors demonstrate that YAP/TAZ is able to control MYL9 by post-translational modifications, placing YAP as a critical factor in regulating matrix-remodeling and invasion through MYL96.
Calvo et al. next posited that YAP/TAZ activation may not be exclusive to CAFs, but may also occur in normal fibroblasts when placed in a cancerous environment6. They found that fibroblasts placed in culture with tumor conditioned media had higher nuclear translocation of YAP, and higher gel contraction (akin to matrix stiffening) comparable to known promoters of pro-contractile function: L-alpha-lysophosphatidic acid (LPA) and transforming growth factor-beta (TGFβ). However, actomyosin inhibition (by blebbistatin) could not be rescued with LPA and TGFβ. Therefore, while soluble factors may activate matrix contraction, a functional cytoskeleton is essential for matrix contraction. Because of the necessary role of the cytoskeleton, the authors tested whether inhibition of RhoA kinase (ROCK), a kinase involved in regulating translocation and structure of the cell by the cytoskeleton, would affect the nuclear localization of YAP6. Inhibition of ROCK decreased YAP nuclearization and decreased the matrix stiffness. Of note, like ROCK inhibition, inhibition of Src also affected the nuclear localization of YAP as well as complex formation with TEAD1 and TEAD4. However, Src modulation of YAP is downstream of cytoskeletal changes in tension since Src inhibition did not affect stress fibers6.
Since activation of YAP in CAFs is connected to actomycin-mediated matrix stiffness, and this activation of YAP expresses MYL9, and expression of MYL9 results in matrix-remodeling towards stiffness, the authors posit that this pathway forms a feed-forward loop6. This loop could lead to constitutive activation of YAP pathway in CAFs, causing a robust response and stabilizing the CAF phenotype. However, it is not known what other mechanisms, as well as regulatory mechanisms of YAP, are involved in this process as well as whether the YAP-ECM tension pathway may play a regulatory role in normal fibroblasts.
References:
1. Boudreau, A., van’t Veer, L. J. & Bissell, M. J. An “elite hacker”: breast tumors exploit the normal microenvironment program to instruct their progression and biological diversity. Cell adhesion & migration 6, 236-248, doi:10.4161/cam.20880 (2012).
2. Levental, K. R. et al. Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell 139, 891-906, doi:10.1016/j.cell.2009.10.027 (2009).
3. Guilluy, C. et al. The Rho GEFs LARG and GEF-H1 regulate the mechanical response to force on integrins. Nature cell biology 13, 722-727, doi:10.1038/ncb2254 (2011).
4. Sawada, Y. et al. Force sensing by mechanical extension of the Src family kinase substrate p130Cas. Cell 127, 1015-1026, doi:10.1016/j.cell.2006.09.044 (2006).
5. Harvey, K. F., Zhang, X. & Thomas, D. M. The Hippo pathway and human cancer. Nature reviews. Cancer 13, 246-257, doi:10.1038/nrc3458 (2013).
6. Calvo, F. et al. Mechanotransduction and YAP-dependent matrix remodelling is required for the generation and maintenance of cancer-associated fibroblasts. Nature cell biology 15, 637-646, doi:10.1038/ncb2756 (2013).

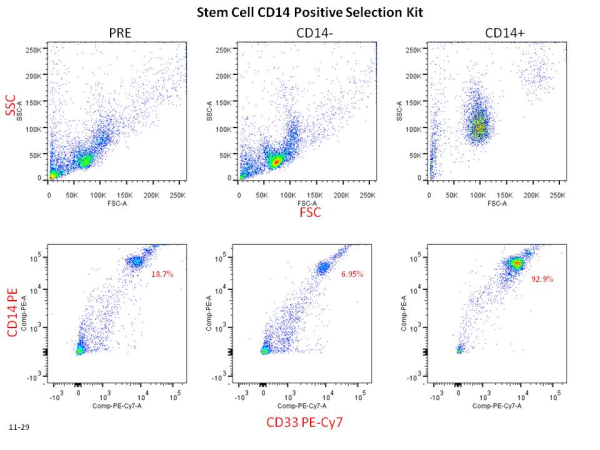

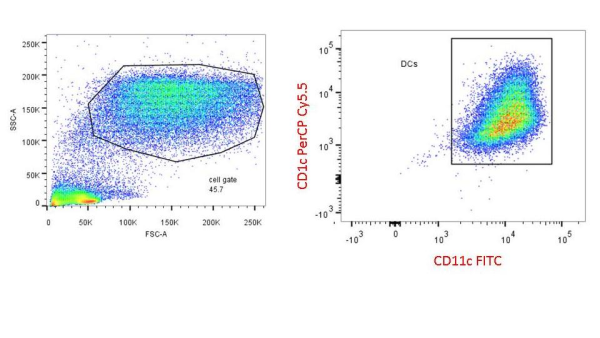



 Colt Egelston is currently a post-doctoral fellow at the Beckman Research Institute of the City of Hope, in Duarte, CA. He received his Ph.D. from Rush University in Chicago and is interested in all things immunology.
Colt Egelston is currently a post-doctoral fellow at the Beckman Research Institute of the City of Hope, in Duarte, CA. He received his Ph.D. from Rush University in Chicago and is interested in all things immunology.
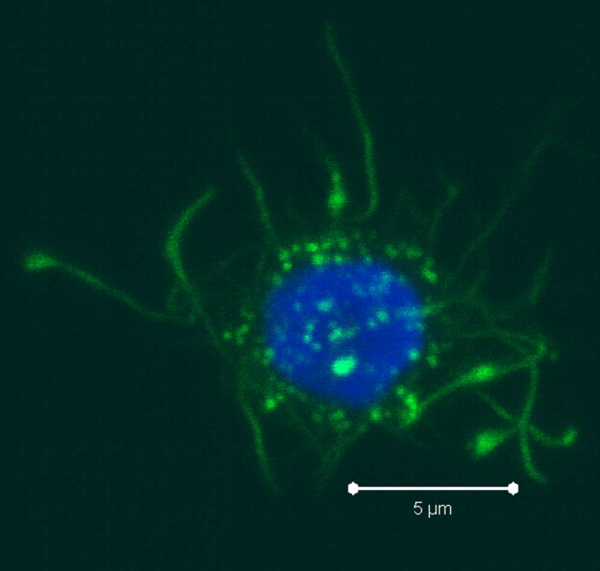
 PBMCs are not just a source of many different circulating immune cell types, but also a source of potential cells that one can generate in vitro. One excellent and long-standing example of this is the generation of dendritic cells (DCs) from monocytes. Monocyte derived DCs (mDCs) are an excellent tool for researchers to do immunological assays requiring a source of professional antigen presenting cells (APCs). While circulating B cells are capable of antigen presentation and T cell activation, they do not offer the robust response that DCs do. The generation of mDCs is a relatively simple protocol that anyone can do with just a source of PBMCs, a few important cytokines, and, of course, some media and incubator space. After this protocol, you will have obtained immature mDCs that can then be matured for use as APCs in your assay.
PBMCs are not just a source of many different circulating immune cell types, but also a source of potential cells that one can generate in vitro. One excellent and long-standing example of this is the generation of dendritic cells (DCs) from monocytes. Monocyte derived DCs (mDCs) are an excellent tool for researchers to do immunological assays requiring a source of professional antigen presenting cells (APCs). While circulating B cells are capable of antigen presentation and T cell activation, they do not offer the robust response that DCs do. The generation of mDCs is a relatively simple protocol that anyone can do with just a source of PBMCs, a few important cytokines, and, of course, some media and incubator space. After this protocol, you will have obtained immature mDCs that can then be matured for use as APCs in your assay. Once the culture period has finished, between 6-8 days, the mDCs can be collected. The exact day is not critical, as long as you remain consistent in the day you pick for your following experiments. To collect the mDCs, gently wash the culture dishes with several streams of media by pipetting up and down. The mDCs, which are currently immature, will be somewhat floating and only loosely adherent. Because of their loose adherence, they require several rounds of gentle pipetting, but do not require cell scraping, EDTA, or trypsin treatment. Note that the culture dishes will still contain some adherent cells. Do not worry about these cells, since these are not the loosely adherent DCs we are interested in.
Once the culture period has finished, between 6-8 days, the mDCs can be collected. The exact day is not critical, as long as you remain consistent in the day you pick for your following experiments. To collect the mDCs, gently wash the culture dishes with several streams of media by pipetting up and down. The mDCs, which are currently immature, will be somewhat floating and only loosely adherent. Because of their loose adherence, they require several rounds of gentle pipetting, but do not require cell scraping, EDTA, or trypsin treatment. Note that the culture dishes will still contain some adherent cells. Do not worry about these cells, since these are not the loosely adherent DCs we are interested in. Colt Egelston is currently a post-doctoral fellow at the Beckman Research Institute of the City of Hope, in Duarte, CA. He received his Ph.D. from Rush University in Chicago and is interested in all things immunology.
Colt Egelston is currently a post-doctoral fellow at the Beckman Research Institute of the City of Hope, in Duarte, CA. He received his Ph.D. from Rush University in Chicago and is interested in all things immunology.
