Whether eating too much table salt in our diet is bad for our health has long been debated. Links have been proposed to several cardiovascular diseases. But, a recent expert committee for the Institute of Medicine concluded that the data do not support such a link [1], keeping the discussion going. Two recent publications in Nature, however, suggest that too much dietary salt might impact our immune system instead and potentially increase the likelihood of autoimmune diseases.
CD4 T lymphocytes can differentiate in specialized subsets that promote or help diverse immune responses. Called T helper (Th) cells, particular subsets are named after prominent cytokines they produce, e.g. IL-17 in the case of Th17 T cells. Th17 cells are important for protection of the body against many bacterial and fungal infections and they are prevalent in the intestinal tissue were they are believed to aid the barrier function of the gut to keep the intestinal bacteria were they belong [2]. However, the “too much of a good thing” proverb applies to lymphocytes too and in the case of Th17 T cells this is exemplified by their pathogenic involvement in several autoimmune diseases. Therefore, the control of cell number and function of Th17 cells requires a delicate balance.
It was known that there is a cross talk between the gut lumen and the Th17 cell response. For example, a few years back it was shown that the frequency of a common bacterium within the gut microbiota could influence the prevalence of Th17 cells in the intestinal tissue [3]. The two new studies demonstrate that table salt (sodium chloride, NaCl) is a surprising new factor on the list to influence the frequency and function of Th17 cells [4-6].
Adding 40 mM NaCl – a level found in the intestinal tissue of animals after feeding of a high salt diet – to in vitro cultures augmented the differentiation of Th17 cells [4, 5]. Similar to in vitro, feeding mice with a high salt diet increased the frequency of Th17 cell in the intestinal tissue, but not in the lymph nodes or the spleen. In both settings (in vitro and in vivo) the resulting Th17 cells were capable of producing large amounts of pro-inflammatory cytokines. By analysis of the mRNA expression, both reports characterized the MAP-kinase p38, NFAT5 (nuclear factor of activated T cells 5) and SGK1 (serum glucocorticoid-regulated kinase-1) as critical molecular players in sensing NaCl and mediating its effect. The elimination of any of these factors from the T cells, either by genetic ablation or by impeding the expression by means of RNA-interference (shRNA), blocked the increased Th17 cell differentiation in the presence of NaCl. Although all three proteins are part of the same pathway, SGK1 appeared to be central in the regulation of the NaCl induced effect. Although this finding is surprising, the results are in line with the known function of SGK1 in sodium transport and homeostasis [7]. SGK1 expression was not only induced by increased NaCl concentrations, but also by the cytokine IL-23, which has a critical role in stabilizing and reinforcing the TH17 phenotype [2]. As NaCl also increased the expression of the IL-23 receptor this established a positive feedback loop that strengthened the Th17 cell differentiation. Importantly, both groups also showed that raising the levels of dietary salt could augment the severity of EAE (experimental autoimmune encephalomyelitis), a mouse model for the autoimmune disease multiple sclerosis [4-6].
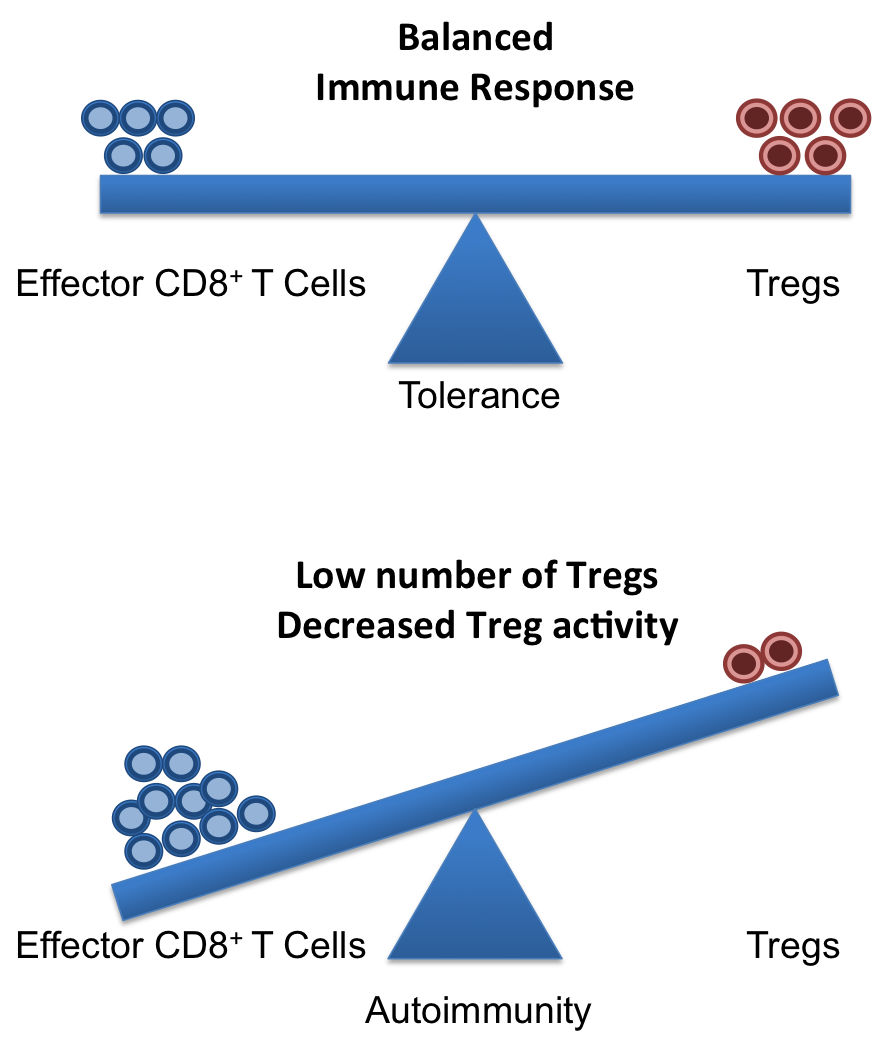
In summary, these reports demonstrate that high levels of salt in the diet could make mice susceptible to a form of autoimmune disease that involves pathogenic Th17 T cells. The data suggest that high concentration of NaCl might be an environmental risk factor for autoimmune diseases. However, it should be pointed out that high concentration of NaCl did not induce autoimmune responses by itself, as the EAE animal model requires the immunization with a know self-antigen. Autoimmunity is a complex interplay of numerous genetic pre-disposing and environmental factors. In this regard these new reports [4, 5] suggest that high dietary salt concentrations might tilt the balance a bit towards autoimmunity in genetically predisposed individuals.
However, the reality will likely be more complicated – as it usually is. For example, it will be critical to show that the correlation between dietary NaCl and Th17 cells is valid also in humans. Furthermore, with this knew knowledge other factors might come to light soon. For example, SGK1 expression is also stimulated by several hormones including endogenous steroids like stress hormones [7], suggesting that the induction of Th17 cell might be augmented by stress as well. Therefore, these intriguing new reports [4, 5] will surely spur now the required research to clarify these points. Till then, going slow on sodium-laden junk food might be generally a justified suggestion.
References:
[1] Strom, Brian (2013). Sodium Intake in Populations: Assessment of Evidence. Washington, DC: The National Academies Press: The Institute of Medicine.
[2] Weaver, C. T., Elson, C. O., Fouser, L. A. & Kolls, J. K. The Th17 pathway and inflammatory diseases of the intestines, lungs, and skin. Annu Rev Pathol 8, 477–512 (2013).
[3] Ivanov, I. I. et al. Induction of Intestinal Th17 Cells by Segmented Filamentous Bacteria. Cell 139, 485–498 (2009).
[4] Kleinewietfeld, M. et al. Sodium chloride drives autoimmune disease by the induction of pathogenic TH17 cells. Nature (2013). doi:10.1038/nature11868.
[5] Wu, C. et al. Induction of pathogenic TH17 cells by inducible salt-sensing kinase SGK1. Nature (2013). doi:10.1038/nature11984.
[6] O’Shea, J. J. & Jones, R. G. Autoimmunity: Rubbing salt in the wound. Nature 496, 437–439 (2013).
[7] Lang, F. & Shumilina, E. Regulation of ion channels by the serum- and glucocorticoid-inducible kinase SGK1. The FASEB Journal 27, 3–12 (2013).